Optimized oligonucleotide synthesis is chemical synthesis of nucleic acids containing additional properties that do not naturally occur in DNA or RNA. Common modifications include fluorophores, quenchers, backbone modifications and unnatural bases. Modified oligonucleotides can be utilized in molecular diagnostics, nucleic acid therapeutics and basic genomics research. Most applications of modified oligonucleotides require increased functionality than unmodified phosphodiester oligonucleotides. Chemical synthesis of modified oligonucleotides is performed with similar phosphoramidite chemistry with purification methods sometimes modified for the modification.
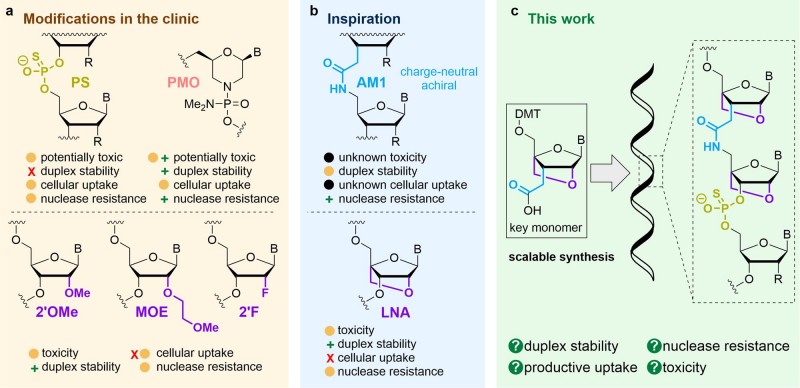 Therapeutic oligonucleotide modifications and our strategy for combining these.1,5
Therapeutic oligonucleotide modifications and our strategy for combining these.1,5
Solid-phase chemical synthesis of modified oligonucleotides can be considered a straightforward extension of standard solid-phase DNA synthesis. Modified oligonucleotides can differ from normal oligonucleotides by containing additional functional groups which alter the chemical and/or biological behavior of DNA or RNA molecules. These functional groups may be linked directly to the internucleotide phosphodiester linkage, or may substitute for the natural nucleosides. Examples of common modifications are fluorescent dyes, backbone modifications or nucleotide analogs.
Table 1 Categories of Oligonucleotide Modifications
| Modification Type | Chemical Nature | Primary Function |
| Backbone modifications | Phosphorothioate, morpholino, PNA | Nuclease resistance, altered charge |
| Sugar modifications | 2'-O-methyl, 2'-fluoro, LNA | Enhanced binding affinity, stability |
| Base modifications | 5-methylcytidine, pseudouridine | Reduced immunogenicity, improved pairing |
| Terminal labels | Fluorophores, biotin, quenchers | Detection, capture, FRET applications |
Native oligonucleotides, also known as regular oligos, have natural phosphodiester linkages between the sugar of either deoxyribose or ribose and the nucleobase. Modified oligonucleotides, also known as chemically modified oligonucleotides, possess a chemical modification that substitutes for part of a natural nucleotide structure. Modifications to oligonucleotides allow for circumvention of many limitations associated with standard oligonucleotides such as instability in serum, cellular uptake, target and protein affinity, and chemical hydrolysis. Modifications can occur in the backbone of the oligonucleotide or sugar, or at the nucleobase itself. Common backbone modifications include phosphorothioates, which confer resistance to nuclease degradation while still allowing RNase H activity against bound RNA targets. Alternative backbones, such as morpholino or peptide nucleic acids, remove the negative charge of the oligonucleotide which can aid in cellular uptake and avoid nucleases entirely at the expense of altering protein binding profile. 2'-Chemical modifications including methoxy, fluoro or methoxyethyl groups can improve affinity and stability without impacting charge. Modified oligonucleotides can have vastly different properties than their unmodified counterparts, often being considered drug-like molecules. Because of this they require different synthesis methods and analytical validation methods than standard oligonucleotides.
Fluorescent and quencher labels can be attached to oligonucleotides using phosphoramidite chemistry for site-specific labelling of nucleic acids at defined sequence locations. Dye molecules can be added precisely where needed: either by directly including dye-phosphoramidites in the solid-phase oligonucleotide synthesis or by chemically attaching dye molecules to the nucleic acid after synthesis, using modified nucleosides introduced during the process. Optimal coupling conditions and detritylation conditions must be determined to allow for dye stability.
Fluorescently labeling oligonucleotides is commonly done with phosphoramidite building blocks that can directly append the dye to the oligonucleotide during automated synthesizers. Dyes like carboxyfluorescein can be appended at the 5'-end or internally using carboxyfluorescein amidites (FAM). Cyanine dyes such as Cy3 and Cy5 are phosphoramidites that emit light further toward the red end of the spectrum. Increased reaction times may be necessary during synthesis to accommodate dye modification due to steric bulk. Modified oligonucleotides must maintain fluorescent integrity during synthesis which includes resistance to the final deprotection step.
Quenchers are often used as fluorescence labels themselves, since they are molecules that lack fluorescence themselves but can absorb energy through Förster resonance energy transfer or static quenching from an adjacent fluorophore. Because of this interaction with fluorophores they can be used to allow detection methods like TaqMan probes or molecular beacons. Other quenchers, referred to as dark quenchers such as Black Hole Quenchers and Dabcyl, can spectrally overlap with a dye used as the donor quencher pair and decrease fluorescence emission only when the donor and acceptor are in close proximity to each other. This forms the basis of signal-off signal-on detection used when they are combined with a fluorophore, and quenching occurs upon hybridization to a target sequence or enzymatic cleavage of the probe. When fluorophores and quenchers are attached to opposite ends or internally on an oligonucleotide, they are referred to as a dual-labelled oligonucleotide.
Modified backbone oligonucleotides are chemically synthesized oligonucleotides in which the normal phosphodiester linkage is replaced with a chemically modified linker. These linkers are typically modified to increase chemical stability, binding strength or enzymatic resistance. Common linkers used include phosphorothioate linkers, phosphoramidate linkers, and morpholinos. Backbone modification chemically alters the nucleic acid polymer, although base-pairing ability is retained. They utilize phosphoramidite chemistry and require different synthesis protocols than DNA synthesis.
 Synthesis cycle for the preparation of PMOs.2,5
Synthesis cycle for the preparation of PMOs.2,5
In phosphorothioate chemistry, sulfur is introduced to the internucleotide linkage (between two phosphate groups) via thiolation of the elongating oligomer. This reaction occurs during solid-phase synthesis. The non-bridging oxygen is replaced with sulfur, forming a diastereomeric phosphorus center. As a result, Rp and Sp phosphorothioates are formed as mixtures. Common sulfurizing agents used after coupling during synthesis include xanthane hydride or Beaucage reagent. These reagents transform phosphite triesters into phosphorothioates. Such linkages show greater resistance to hydrolysis by nucleases and greater serum protein binding compared to normal phosphate oligomers. These effects lead to improved half-life for oligonucleotide therapeutics, from minutes to days. On the downside, phosphorothioate linkages introduce stereochemistry to oligonucleotides because each phosphorothioate linkage can occur as a pair of epimers, which may possess unique biological characteristics. Traditional synthesis methods will result in a stereorandom mixture with multiple diastereomers. Methods which produce pure Rp or Sp configurations have been developed using chiral auxiliaries or oxazaphospholidine monomers, but require modified coupling reactions and longer reaction times to reach completion.
Control of chirality for phosphorothioates means controlling the Rp or Sp configuration of the sulfur modification. The phosphorothioate linkage behaves as a stereocenter and can exist in two configurations, Rp or Sp. Compounds containing both configurations will exist as diastereomers of each other. Dependent on which stereoisomer is placed at each linkage along an oligonucleotide the final product can have differing therapeutic characteristics. If synthesized stereorandomly, each synthesis will result in random placement of Rp and Sp linkages throughout the oligonucleotide. Complete stereochemical control of phosphorothioates can be achieved through the use of chiral phosphoramidite monomers. These monomers then couple with controlled stereochemistry at the sulfur. The chiral auxiliary is then removed while maintaining configuration. To ensure batch-to-batch consistency of the stereoisomer distribution, the product should be well characterized by methods that can distinguish between Rp and Sp linkages. Control of chirality at the phosphorothioate linkages can alter sensitivity to nucleases and protein binding; however, batch-to-batch consistency can be maintained by keeping the stereorandom incorporation consistent from batch to batch instead of controlling chirality completely.
Base-modified oligonucleotides are essentially nucleosides that have been tweaked with unconventional nucleobases. These bases are modified to have altered Watson-Crick pairing properties, base stacking, or nucleobase reactivity while still preserving the DNA backbone. Specialized properties include high affinity, mismatched base pairing, and different or novel chemical reactivity than A, G, C, or T. Applications include site-directed mutagenesis, DNA footprinting, and antisense drugs. Modified phosphoramidites allow for chemical synthesis on an automated DNA synthesizer.
Table 2 Classes of Modified Nucleobases and Their Functional Properties
| Modified Base | Structural Alteration | Functional Effect | Primary Application |
| 5-Bromouracil | Halogen substitution at C5 | Increased stability; photoreactivity | Crosslinking studies; mutagenesis |
| Inosine | Hypoxanthine base | Degenerate pairing (A/C/G/T) | Universal probe design |
| 2-Aminopurine | Amino group at C2 | Fluorescent properties | Structural dynamics probing |
| 5-Methylcytosine | Methyl group at C5 | Enhanced stacking; epigenetic mimic | Methylation analysis |
| Locked nucleic acids | Bridged 2'-O,4'-C linkage | Conformational restriction | Affinity enhancement |
Modified nucleobases are introduced by using appropriately modified phosphoramidite monomers containing the modified heterocycle instead of the natural base. They are added to a growing chain using automated solid-phase synthesis. Conditions are optimized for coupling of modified nucleotides, which have different steric and electronic demands than natural nucleotides. Some modifications require longer coupling times or alternate activators to obtain adequate coupling efficiencies. After chain assembly, modified oligonucleotides are deprotected under conditions that do not harm the modification. For example, some modified bases are sensitive to aqueous ammonia allowing for milder deprotection conditions. Site-specific modified bases can be placed internally or at the oligonucleotide terminus to optimize oligonucleotide properties. Another type of modification incorporates universal bases, which can bond with multiple natural bases, and photoreactive crosslinking groups that link to proteins when exposed to UV light.
Changes to bases can affect the thermodynamics of hybridization and stability of a duplex by changing hydrogen bond donors and acceptors, base stacking, and backbone structure. Stabilizing additions can increase melting temperature, Tm, by increasing stacking interactions. Examples include adding hydrophobic groups to bases. Destabilizing changes can also increase specificity to a target sequence by changing hydrogen bond donors and acceptors. These changes can decrease Tm. Modified bases can be universal and pair with more than one natural base. Degenerate primers can then be created to amplify various sequences using polymerase chain reaction. Destabilizing mismatches can be used to detect single nucleotide polymorphisms, SNPs, during genotyping. Changes in one base can create enough change in either hybridization rates or duplex stability that they can be detected. Modifications to bases can affect hybridization, so they have to be changed with caution. Targets have to be recognized without affecting the stability needed for the probe to function correctly or be processed by enzymes.
Chemically modified oligonucleotide synthesis can be challenging. Synthesis becomes more challenging as modifications become larger and as the length of the oligonucleotide increases. While the standard phosphoramidite method generally works well for modified oligo synthesis, unusual nucleoside phosphoramidites often struggle, leading to poor coupling and unwanted side effects that hurt the final product's quality. Given that synthesis happens one step at a time, a single misstep in adding a nucleotide will affect the next round. Once long sequences are made, this can quickly result in little to no full-length product. These effects are even more pronounced when synthesizing modified oligonucleotides because modified building blocks can have quite different reactivity from natural nucleoside phosphoramidites.
Coupling inefficiency is often due to the bulk caused by attachments like fluorophores, substitutions on the backbone, or linked ligands. The bulkier the modified phosphoramidite monomer, the slower the coupling step because its incoming amidite nucleophile is sterically hindered from attacking the chain-terminal hydroxyl group. To ensure proper stepwise yield longer coupling times may be necessary. Modified nucleotides often have coupling conditions that need to be changed from those used for natural nucleotides. Conditions can need to be twice as long or three times as long as natural nucleotide coupling steps and higher levels of activator may be needed to force the condensation reaction to completion. Some modifications can reduce the electrophilicity of the phosphorous atom of the phosphoramidite making coupling even slower. Inefficient coupling leads to shorter sequences, deletion variants, and lowers your crude yield. Lower crude yields can make it difficult to purify your product and you may have to resynthesize your compound to get enough product for your desired application.
Modified oligonucleotide synthesis is associated with high levels of side products due to incomplete coupling reactions, depurination, reactions at the modified base or backbone, or reactions caused by structural crowding. Short chains can be formed when incomplete coupling reactions are followed by capping of the unextended chains. In these instances, uncapped chains can accept another nucleotide causing deletion products to accumulate. Structure formed by steric bulk of the modification can block access of reagents to the 3'-terminus of a growing oligonucleotide chain allowing mispriming or non-template reactions. Modified bases and linkages can have lower acid stability than the natural nucleotides. Modified oligonucleotides are therefore more prone to experience depurination during the acid detritylation step or during chemical degradation in the cleavage/deprotection reaction. Side products from incomplete reactions can be difficult to remove during purification because they can co-elute with full-length product during reversed-phase chromatography or gel purification. More extensive purification by high-performance liquid chromatography or gel purification may be required to produce homogeneous material.
The analytical purification and quality control processes for modified oligos are much more complex than for regular DNA. Modified oligos that contain fluorescence dyes, backbone modifications, or chemically conjugated entities, tend to result in a product mixture of failure sequences, truncated deprotection species and stereoisomers that make it difficult to isolate a pure species. Purification of modified oligos must separate failure sequences from full-length sequences while maintaining structural integrity of labile modifications. Therefore, many types of chromatography including reversed-phase chromatography, ion-exchange chromatography and size exclusion chromatography are used. Identity, integrity of the modification, purity, and consistency must be monitored and confirmed using appropriate analytical techniques including mass spectrometry to guarantee quality control of the finished product. The methods used depend on the modification, application of use and desired purity. If the oligo is intended for use in therapeutic applications stringent characterization is required. However, if the oligos are only used for research purposes they require less characterization.
Modified oligonucleotides can be characterized using many standard techniques. The methods chosen will depend on the chemical modifications present on the oligonucleotide. Modifications to the backbone will often not absorb UV light. For example, phosphorothioate modifications will require changing the detection system or MS verification. Modified full length product can often be separated from impurities using ion-pair reversed-phase HPLC. Ion pair reversed phase liquid chromatography methods can easily be MS compatible by choosing the correct mobile phase. Longer oligonucleotides with more modification will require additional methods to account for all deletion, addition, and chemically modified impurities. The chromatography method chosen must separate all the impurities but also account for time constraints. For example, preparative chromatography would focus on yield, but analytical chromatography methods would be able to focus on detection limits.
The chromatographic separation of modified oligonucleotides needs to be optimized depending on the stationary and mobile phase chemistries used to resolve the target product from closely related synthetic byproducts. Reversed-phase chromatography relies on the separation of complete full-length oligonucleotides (still containing capping reagents at the termini) and shorter failure sequences. Resolution may be lessened due to secondary structure interactions when attempting to resolve longer oligonucleotides. Ion exchange chromatography offers different selectivity in separating modified oligonucleotides from side products that contain different numbers of phosphate or phosphorothioate groups. Unique purification considerations should be taken into account when purifying oligonucleotides modified with bulk groups like fluorophores or streptavidin.
Use of mass spectrometry (MS) is recommended to confirm the identity and integrity of any synthesized modified oligonucleotide. MS can provide molecular weight data that should match the expected weight of the synthesized oligonucleotide, thus confirming that the correct compound was synthesized. Mass spectrometry is especially important for modified oligonucleotides because it can confirm that modifications such as fluorescent tags added to the ends of the oligonucleotide, backbone modifications like phosphorothioate groups or nucleotide substitutions are present and intact after synthesis and purification. Multiply charged ions formed using electrospray ionization can be used to determine the mass of long oligonucleotides. Matrix-assisted laser desorption can be used for rapid mass determination of shorter oligonucleotides. It is important to analyze the distribution of charge and possible formation of adducts when analyzing the MS data. Any molecules present that have a molecular weight different from that of the expected oligonucleotide could be due to incomplete detritylation, side reactions that occurred during synthesis, or degradation products that can affect activity. MS can be used to document compliance with regulatory filings for therapeutic oligonucleotides.
Modified oligonucleotides are used for many purposes in basic research as well as clinical diagnostic and therapeutic applications. Many natural forms of nucleic acids lack desirable chemical and physical properties for certain applications. For example, unmodified oligonucleotides are rapidly metabolized in cells, they may not enter cells efficiently, and they may lack sufficient binding affinity for their targets. Modifications such as backbone modifications, sugar modifications, and conjugation to various entities can help to solve these problems. Modified oligonucleotides can be used as molecular probes for detecting nucleic acids or proteins. Modified oligonucleotides are used as therapeutic agents to target disease areas that were previously considered undruggable. Modified oligonucleotides are also used for gene editing. Modifications useful for molecular probes can include fluorescence-quenching properties and photostability. Modifications useful for therapeutics can include resistance to nucleases and altered pharmacokinetics. Modifications useful for gene editing can include protein compatibility.
Table 3 Application-Specific Modification Requirements for Oligonucleotides
| Application Domain | Critical Modification Types | Functional Objectives |
| Molecular diagnostics | Fluorescent dyes, quenchers | Signal generation and detection sensitivity |
| RNA therapeutics | Backbone modifications, sugar modifications | Metabolic stability and reduced immunogenicity |
| Gene editing systems | Chemical stabilization of guide RNAs | Structural integrity in cellular environments |
| Targeted therapeutics | Conjugation chemistries | Cell-specific delivery and enhanced pharmacokinetics |
Chemically modified oligonucleotides can be used to build advanced diagnostic tools because of their versatility and sensitivity. Molecular beacons and TaqMan probes allow the sensitive detection of target nucleic acids by labeling with reporter dyes and quenchers. They remain fluorescenceless as long as the reporter and quencher are adjacent. However, once the oligonucleotide finds its target, it hybridizes and the fluorescent signal can be detected. Modifications need to be photostable and have emission wavelengths that can be distinguished from one another while still binding to their complementary sequence of interest. Capture probes can be anchored onto a surface via biotin or another affinity tag allowing for the purification and concentration of nucleic acid targets from biological samples for clinical diagnostics such as infectious disease or SNP analyses. Backbone modifications allow probes to be more stable during PCR and heating steps.
Chemical modification is required for RNA therapeutics, such as antisense oligonucleotides, siRNA and mRNA, to be effective drugs. Common modifications include alterations to the backbone such as phosphorothioate that help the RNA avoid degradation by serum nucleases and allow interaction with RNA processing proteins. Modifications at the ribose 2'-position, such as 2'-fluoro or 2'-methoxy groups can increase binding to target RNAs and serum stability. Modifications to nucleobases such as pseudouridine can also alleviate immune stimulation caused by the therapeutic when administered. Chemical modifications can increase the half-life of oligonucleotides in circulation from seconds to hours or days. With an adequate half-life, the therapeutic can distribute into tissues to alter expression of its target gene by degradation, alteration of splicing or prevention of translation.
Modified oligonucleotides are also used for gene editing applications. They can be employed as parts of engineered nuclease systems to create site-specific changes to the genome. For example, sgRNAs used with CRISPR-Cas systems need to be chemically modified to be stable inside of cells. Cas proteins induce a double strand break at the desired genomic location after the guide RNA-localizes to that specific site. The modifications prevent degradation of the guide RNA before it can bind to the endonuclease. Modified oligonucleotides can also be used as donor DNA or repair oligos to leverage homology directed repair to fix the edit. Linker modifications allow these oligonucleotides to evade degradation inside of the cell. Custom chemical synthesis of these oligonucleotides allows for site specific changes to be made to correct genetic disease.
An aptamer is an oligonucleotide, single-stranded DNA or RNA molecule that binds to a specific target molecule. They are selected from a large pool of nucleic acid sequences in an iterative process known as SELEX (Systematic Evolution of Ligands by Exponential enrichment). Aptamers fold into complex shapes, enabling them to bind targets with high affinity and specificity. Aptamers can serve as mimetics to antibodies for diagnostic and therapeutic use. Chemical modification allows aptamers to become more metabolically stable for in vivo use. Modifications to the two-prime position of the ribose sugar backbone with fluoro, amino or methoxy groups can allow for resistance to nuclease digestion. Additionally, modification of the backbone can increase metabolic stability. Modification of the termini with high molecular weight polymers can increase circulating half-life by limiting renal clearance. Aptamers can target specific cells. They have been used as monotherapeutics that work by blocking the target receptor and can be conjugated to drugs, imaging reagents, and other payloads.
Modified oligonucleotide synthesis requires far more than standard solid-phase chemistry. The incorporation of fluorescent labels, backbone modifications, and base analogs introduces additional chemical complexity that directly impacts coupling efficiency, purification strategy, and final product performance. Our technical capabilities in modified oligonucleotide synthesis include:
By integrating optimized chemistry, purification expertise, and rigorous quality control, we support applications ranging from advanced research assays to regulated diagnostic and therapeutic programs.
Highly modified oligonucleotides often involve design trade-offs between stability, performance, yield, and scalability. Early technical evaluation can help prevent synthesis inefficiencies, purity challenges, and downstream performance issues. If your project involves:
our scientific team can review your specifications and provide guidance on synthesis strategy, purification approach, and expected performance considerations. Contact us to discuss your modified oligonucleotide project and explore the most suitable synthesis solution for your application.
References
It is an oligonucleotide containing chemical changes to bases, sugars, backbones, or termini.
Labels are added during synthesis using modified phosphoramidites or via post-synthesis conjugation.
Yes, complex modifications often lower yield and require optimized synthesis conditions.
Modifications increase side products and sequence heterogeneity that must be removed.
Diagnostics, RNA therapeutics, antisense, and gene editing applications.

